A simple methodology for RNA isolation from bacteria by integration of formamide extraction and chitosan-modified silica purification

RNA isolation from bacteria is technically difficult due to the RNA characteristic of labile and vulnerable degradation. Many reagents were explored for cellular lysis and complete inhibition of RNase. However, the available methods for RNA isolation are either of low efficiency or time-consuming. Here, we developed a rapid and accessible protocol for RNA isolation that combined a simplified cell lysis and RNA release by formamide-based solution and RNA purification by chitosan-modified silica membrane for the first time. With this method, we obtained about ~ 28 μg of total RNA from 10 8 Escherichia coli cells. The entire procedure can be done within 15 min without redundant pipetting steps. The purity of extracted RNA was comparable to that of commercial kits, but the cost was much lower. Furthermore, the yielded RNA was successfully used in downstream enzymatic reactions, such as reverse transcription and quantitative real-time PCR. This new method would be of benefit for an extensive range of gene expression analyses in bacterial organisms.
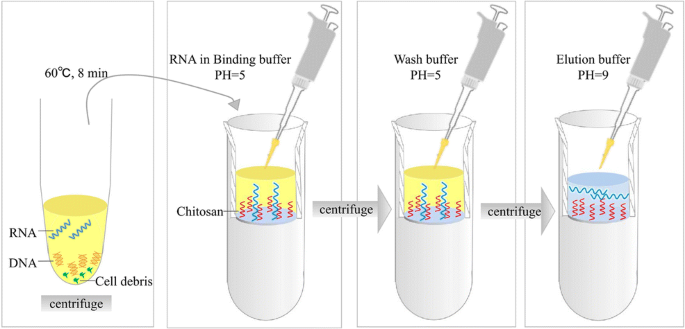
Graphical abstract
This is a preview of subscription content, log in via an institution to check access.
Access this article
Subscribe and save
Springer+ Basic
€32.70 /Month
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
Buy Now
Price includes VAT (France)
Instant access to the full article PDF.
Rent this article via DeepDyve
Similar content being viewed by others
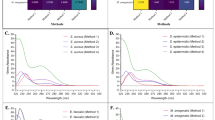

Effective RNA isolation method for gram-positive and acid-fast bacteria: metamorphosed from conventional RNA isolation techniques
Article 07 August 2024

Extraction of Total RNA from Yersinia pestis
Chapter © 2018

The Single-Step Method of RNA Purification Applied to Leptospira
Chapter © 2020
References
- Higo S, Higo M, Tanifuji S. Specific dissociation of pea 25-S rRNA by hot-phenol treatment. Biochim Biophys Acta. 1971;246(3):499–506. https://doi.org/10.1016/0005-2787(71)90787-8. ArticleCASPubMedGoogle Scholar
- Chirgwin JM, Przybyla AE, MacDonald RJ, Rutter WJ. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry. 1979;18(24):5294–9. https://doi.org/10.1021/bi00591a005. ArticleCASPubMedGoogle Scholar
- Glisin V, Crkvenjakov R, Byus C. Ribonucleic acid isolated by cesium chloride centrifugation. Biochemistry. 1974;13(12):2633–7. https://doi.org/10.1021/bi00709a025. ArticleCASPubMedGoogle Scholar
- Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987;162(1):156–9. https://doi.org/10.1006/abio.1987.9999. ArticleCASPubMedGoogle Scholar
- Chomczynski P. Solubilization in formamide protects RNA from degradation. Nucleic Acids Res. 1992;20(14):3791–2. https://doi.org/10.1093/nar/20.14.3791. ArticleCASPubMedPubMed CentralGoogle Scholar
- Stead MB, Agrawal A, Bowden KE, Nasir R, Mohanty BK, Meagher RB, et al. RNAsnap: a rapid, quantitative and inexpensive, method for isolating total RNA from bacteria. Nucleic Acids Res. 2012;40(20):e156. https://doi.org/10.1093/nar/gks680. ArticleCASPubMedPubMed CentralGoogle Scholar
- Shedlovskiy D, Shcherbik N, Pestov DG. One-step hot formamide extraction of RNA from Saccharomyces cerevisiae. RNA Biol. 2017;14(12):1722–6. https://doi.org/10.1080/15476286.2017.1345417. ArticlePubMedPubMed CentralGoogle Scholar
- Bowrin V, Rouse-Miller J, Sutton F, Sirju-Charran G. Formamide-based RNA isolation at above zero temperatures from high starch cassava tubers. Phytochem Anal. 2013;24(1):93–6. https://doi.org/10.1002/pca.2390. ArticleCASPubMedGoogle Scholar
- Cao W, Easley CJ, Ferrance JP, Landers JP. Chitosan as a polymer for pH-induced DNA capture in a totally aqueous system. Anal Chem. 2006;78(20):7222–8. https://doi.org/10.1021/ac060391l. ArticleCASPubMedGoogle Scholar
- Jiang C, Xu S, Zhang S, Jia L. Chitosan functionalized magnetic particle-assisted detection of genetically modified soybeans based on polymerase chain reaction and capillary electrophoresis. Anal Biochem. 2012;420(1):20–5. https://doi.org/10.1016/j.ab.2011.09.004. ArticleCASPubMedGoogle Scholar
- Reedy CR, Price CW, Sniegowski J, Ferrance JP, Begley M, Landers JP. Solid phase extraction of DNA from biological samples in a post-based, high surface area poly(methyl methacrylate) (PMMA) microdevice. Lab Chip. 2011;11(9):1603–11. https://doi.org/10.1039/c0lc00597e. ArticleCASPubMedPubMed CentralGoogle Scholar
- Byrnes SA, Bishop JD, Lafleur L, Buser JR, Lutz B, Yager P. One-step purification and concentration of DNA in porous membranes for point-of-care applications. Lab Chip. 2015;15(12):2647–59. https://doi.org/10.1039/c5lc00317b. ArticleCASPubMedGoogle Scholar
- Gan W, Gu Y, Han J, Li CX, Sun J, Liu P. Chitosan-modified filter paper for nucleic acid extraction and “in situ PCR” on a thermoplastic microchip. Anal Chem. 2017;89(6):3568–75. https://doi.org/10.1021/acs.analchem.6b04882. ArticleCASPubMedGoogle Scholar
- Wang X, Yan C, Wang X, Zhao X, Shi C, Ma C. Integrated silica membrane-based nucleic acid purification, amplification, and visualization platform for low-cost, rapid detection of foodborne pathogens. Anal Bioanal Chem. 2020;412(25):6927–38. https://doi.org/10.1007/s00216-020-02823-1. ArticleCASPubMedGoogle Scholar
- Li Y, Hoffmann J, Li Y, Stephano F, Bruchhaus I, Fink C, et al. Octopamine controls starvation resistance, life span and metabolic traits in Drosophila. Sci Rep. 2016;6:35359. https://doi.org/10.1038/srep35359. ArticleCASPubMedPubMed CentralGoogle Scholar
- Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29(9):e45. https://doi.org/10.1093/nar/29.9.e45. ArticleCASPubMedPubMed CentralGoogle Scholar
- Woldringh CL, van Iterson W. Effects of treatment with sodium dodecyl sulfate on the ultrastructure of Escherichia coli. J Bacteriol. 1972;111(3):801–13. https://doi.org/10.1128/JB.111.3.801-813.1972. ArticleCASPubMedPubMed CentralGoogle Scholar
- Fabre AL, Colotte M, Luis A, Tuffet S, Bonnet J. An efficient method for long-term room temperature storage of RNA. Eur J Hum Genet. 2014;22(3):379–85. https://doi.org/10.1038/ejhg.2013.145. ArticleCASPubMedGoogle Scholar
- Sajomsang W, Gonil P, Tantayanon S. Antibacterial activity of quaternary ammonium chitosan containing mono or disaccharide moieties: preparation and characterization. Int J Biol Macromol. 2009;44(5):419–27. https://doi.org/10.1016/j.ijbiomac.2009.03.003. ArticleCASPubMedGoogle Scholar
- Ross L, Vavro CL, Kehne SL, McClernon DR, St Clair M. Substitution of a commercially available, RNA extraction procedure in an HIV-1 genotyping system improves sensitivity and allows reduced sample volume. J Virol Methods. 2001;96(1):1–4. https://doi.org/10.1016/s0166-0934(00)00275-5. ArticleCASPubMedGoogle Scholar
- Jothikumar N, Sobsey MD, Cromeans TL. Development of an RNA extraction protocol for detection of waterborne viruses by reverse transcriptase quantitative PCR (RT-qPCR). J Virol Methods. 2010;169(1):8–12. https://doi.org/10.1016/j.jviromet.2010.06.005. ArticleCASPubMedGoogle Scholar
- Wang L, Stegemann JP. Extraction of high quality RNA from polysaccharide matrices using cetyltrimethylammonium bromide. Biomaterials. 2010;31(7):1612–8. https://doi.org/10.1016/j.biomaterials.2009.11.024. ArticleCASPubMedGoogle Scholar
- Gomes SL, Gober JW, Shapiro L. Expression of the Caulobacter heat shock gene DnaK is developmentally controlled during growth at normal temperatures. J Bacteriol. 1990;172(6):3051–9. https://doi.org/10.1128/jb.172.6.3051-3059.1990. ArticleCASPubMedPubMed CentralGoogle Scholar
- Narberhaus F, Weiglhofer W, Fischer HM, Hennecke H. Identification of the Bradyrhizobium japonicum DegP gene as part of an operon containing small heat-shock protein genes. Arch Microbiol. 1998;169(2):89–97. https://doi.org/10.1007/s002030050547. ArticleCASPubMedGoogle Scholar
Funding
This work was financially supported by the National Natural Science Foundation of China (81801264), National Key Research and Development Programs of China (2018YFE0113300) and the Key Project of Natural Science Foundation of Shandong Province (ZR2020KH030).
Author information
- Xiaoli Zhao and Yong Li contributed equally to this work.
Authors and Affiliations
- Shandong Provincial Key Laboratory of Biochemical Engineering, Qingdao Nucleic Acid Rapid Detection Engineering Research Center, College of Chemistry and Molecular Engineering, College of Marine Science and Biological Engineering, Qingdao University of Science and Technology, Qingdao, 266042, Shandong, China Xiaoli Zhao, Yong Li, Yake Duan & Cuiping Ma
- Biology Department, College of Science, UAE University, Al Ain, 15551, UAE Amr Amin
- Biology Department, School of Sciences and Humanities, Nazarbayev University, Nur-Sultan, Kazakhstan, 010000 Yingqiu Xie
- Qingdao Nucleic Acid Rapid Testing International Science and Technology Cooperation Base, College of Life Sciences, Department of Pathogenic Biology, School of Basic Medicine, and Department of Clinical Laboratory, The Affiliated Hospital of Qingdao University, Qingdao University, Qingdao, 266071, Shandong, China Chao Shi
- Xiaoli Zhao







